What Are Ligands for E3 Ligase?
Ligands for E3 ligase are small-molecule or peptide-based recognition elements used to recruit E3 ubiquitin ligases in targeted protein degradation research. In a typical PROTAC molecule, an E3 ligase ligand is connected to a target-binding ligand through a linker, creating a bifunctional degrader that can bring the protein of interest close to an E3 ligase system. This induced proximity supports ubiquitin transfer to the target protein and enables downstream degradation through the ubiquitin–proteasome pathway.
For researchers developing PROTACs, molecular glue-inspired degraders, or related induced-proximity tools, the E3 ligase ligand is not only a binding motif. It is a strategic design component that affects ternary complex geometry, degrader selectivity, linker attachment, synthetic route planning, and project scalability. BOC Sciences provides Ligand for E3 Ligase products and related technical support to help discovery teams select suitable E3 ligase recruitment motifs for different protein degradation workflows.
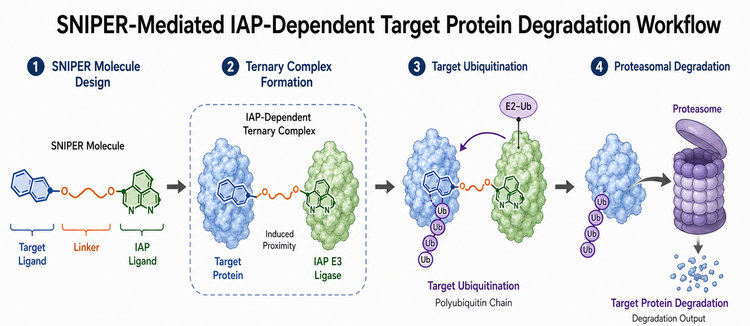 Fig.1 SNIPER-Mediated IAP-Dependent Target Protein Degradation Workflow (BOC Sciences Original).
Fig.1 SNIPER-Mediated IAP-Dependent Target Protein Degradation Workflow (BOC Sciences Original).
How E3 Ligase Ligands Recruit Ubiquitin Ligase Systems?
E3 ligase ligands recruit ubiquitin ligase systems by binding to a ligandable pocket, groove, or surface region of the ligase or substrate-recognition component. In PROTAC research, this recruitment event creates one side of the ternary complex. The linker and target ligand then help position the protein of interest near the recruited E3 ligase machinery. When the assembly is productive, ubiquitin can be transferred to accessible lysine residues on the target protein.
- Ligase recognition: The ligand must bind a chemically tractable region of the E3 ligase system without preventing recruitment into the degrader complex.
- Substrate positioning: The ligand helps orient the E3 ligase so that the target protein can be presented in a ubiquitination-compatible arrangement.
- Linker compatibility: A usable ligand needs an attachment position that tolerates modification while preserving the desired E3 binding mode.
- Research utility: Different E3 ligase ligands allow researchers to compare recruitment strategies and identify degrader designs that fit the biological question.
Why E3 Ligase Ligand Selection Matters?
E3 ligase ligand selection matters because PROTAC activity depends on more than two independent binding events. The degrader must recruit a target protein and an E3 ligase into a productive ternary complex with a geometry that supports ubiquitin transfer. The E3 ligase ligand defines which ligase system is recruited and how that ligase is presented in relation to the target-binding side of the molecule.
Influence on Ternary Complex Formation
The same target ligand can show different degradation behavior when paired with different E3 ligase ligands. This is because each ligase system has its own surface topology, ligand-binding orientation, and protein interaction environment. A productive ternary complex requires the E3 ligase ligand, linker, and target ligand to work together. If the selected ligand positions the ligase poorly, the degrader may bind both partners but fail to support efficient ubiquitination.
Impact on Target Ubiquitination and Degradation Efficiency
After a ternary complex forms, the target protein must be positioned so that ubiquitination can occur on accessible residues. The E3 ligase ligand can affect this step by controlling the orientation and proximity of the recruited ligase. In practical degrader optimization, researchers often compare E3 ligase ligand classes, linker attachment points, and linker lengths to determine which combination produces the most informative degradation profile for the target system.
For research teams building degrader candidates, ligand selection is therefore both a chemistry decision and a mechanistic design decision. BOC Sciences supports ligand design for E3 ligase projects where customers need to evaluate ligand type, exit vector, linker compatibility, and early optimization plans.
Structural Features of E3 Ligase Ligands
E3 ligase ligands vary widely in size, polarity, three-dimensional shape, and available modification sites. Useful ligands must combine adequate E3 recognition with practical chemistry for degrader assembly. In many projects, the most important structural questions are: where does the ligand bind, which part of the ligand can be modified, and how does that modification affect recruitment of the E3 ligase system?
Recognition Motifs for E3 Ligase Binding
The recognition motif is the ligand region that engages the E3 ligase or substrate receptor. It may include hydrogen-bonding groups, hydrophobic surfaces, heteroatoms, stereochemical elements, or ring systems that complement the binding site. For a ligand to be useful in PROTAC research, the recognition motif should remain intact after linker attachment. Even small structural changes near the binding pharmacophore can alter ligase engagement, ternary complex stability, or selectivity across degrader analogs.
Common recognition-related features include:
- Hydrogen-Bonding Motifs
- Hydrophobic Contact Regions
- Stereochemical Control
- Heteroatom Placement
- Ligase-Binding Rings
- Exit Vector Tolerance
Functional Handles for Linker Attachment
Functional handles enable the E3 ligase ligand to be connected to a linker or to a preassembled linker–target ligand fragment. These handles must be positioned so that the binding motif is preserved while the linker points toward a productive space. Common attachment chemistry may involve amines, carboxylic acids, alcohols, alkynes, azides, halides, or activated groups. The correct handle depends on the ligand scaffold, linker type, and synthetic route.
Frequently evaluated handle types include:
- Amine Handles
- Carboxylic Acid Handles
- Alcohol Handles
- Azide Handles
- Alkyne Handles
- Activated Groups
Common E3 Ligase Ligand Classes
Several E3 ligase ligand classes are frequently used in PROTAC and targeted protein degradation research. Each class offers a different combination of ligase recruitment profile, structural requirements, linker attachment options, and research utility. Selection should be based on the target protein, assay format, project stage, and desired comparison strategy rather than on one ligand class alone.
CRBN Ligands for PROTAC Construction
CRBN ligands are among the most widely used E3 ligase recruitment motifs in PROTAC research. They are valued because several ligand scaffolds and modification positions have been explored extensively in degrader construction. These ligands are often used when researchers want to build compact degraders, compare linker positions, or create matched analogs for structure–activity relationship studies. BOC Sciences provides CRBN-oriented E3 ligase ligand options that can support early screening and focused degrader optimization.
They are commonly considered for:
- CRBN Recruitment
- PROTAC Construction
- Linker Attachment
- Degrader SAR
VHL Ligands for Bifunctional Degrader Design
VHL ligands provide another important route for E3 ligase recruitment in bifunctional degrader design. They are often selected to compare how a target protein responds to a different ligase system, binding orientation, and ternary complex interface. VHL ligand selection usually requires attention to stereochemistry, linker exit vector, and compatibility with the target ligand. Related background can be explored through resources such as Progress in the Development of E3 Ligase VHL Ligand.
Key design considerations include:
- Stereochemical Fit
- Exit Vector Control
- Linker Compatibility
- Ternary Complex Geometry
IAP and MDM2 Ligands for Alternative Recruitment Strategies
IAP and MDM2 ligands can provide alternative recruitment strategies when a project requires broader E3 ligase comparison. These ligand classes may help researchers test whether a target protein is more effectively positioned by a different ligase system. Because each E3 system brings different expression context, binding orientation, and interaction surfaces, these ligands are typically evaluated through carefully planned screening workflows rather than assumed to be interchangeable.
Emerging E3 Ligase Ligands for Expanded TPD Research
Beyond commonly used ligase systems, emerging E3 ligase ligands are expanding the research space for targeted protein degradation. These ligands may support exploration of additional tissue contexts, subcellular environments, or protein interaction networks. Emerging ligand systems often require more validation because binding, ternary complex behavior, and degradation outcomes may be less predictable. BOC Sciences can assist customers who need to assess ligand options for exploratory E3 recruitment strategies.
Mechanistic Role of E3 Ligase Ligands in PROTAC Research
The mechanistic role of an E3 ligase ligand is to recruit and position a ubiquitin ligase system within a degrader-induced complex. The ligand affects the formation, orientation, and functional outcome of the ternary complex. Its contribution becomes especially important when researchers compare degraders that share the same target ligand but differ in E3 recruitment motif or linker attachment site.
E3 Ligase Engagement and Substrate Positioning
The E3 ligase ligand first engages the selected ligase system and creates the recruitment side of the degrader. Once the target ligand binds the protein of interest, the degrader can bring both proteins into proximity. Productive substrate positioning depends on the orientation created by the E3 ligand, linker, and target ligand together.
Linker-Mediated Assembly of the Ternary Complex
The linker connects the E3 ligand to the target-binding ligand and determines how the two binding events are spatially coupled. A suitable linker can help the ligand-bound E3 ligase and target protein form a ternary complex. BOC Sciences offers related PROTAC linker products for teams evaluating linker length, flexibility, and attachment chemistry.
Ubiquitin Transfer and Target Protein Processing
A productive ternary complex places the target protein in a position that can support ubiquitin transfer. This step depends on accessible residues on the target protein, the recruited E3 ligase system, and the geometry of the degrader assembly. Researchers may combine chemical design with protein ubiquitination services to evaluate whether a selected recruitment strategy supports the intended mechanism.
Ligand-Driven Selectivity Across Degrader Designs
E3 ligase ligand choice can influence degrader selectivity because different ligase systems create different protein–protein interfaces. A ligand that supports cooperative ternary complex formation may produce a distinct degradation profile from another ligand class. This is why matched comparison of E3 ligase ligands can be highly informative in degrader design.
How to Select Ligands for E3 Ligase?
Selecting ligands for E3 ligase requires a balance of biological relevance, chemical feasibility, and project objective. A strong selection process begins with the E3 ligase type, available ligand classes, linker attachment position, and expected compatibility with the target ligand. It then considers whether the ligand will be used for early screening, focused analog development, or a more specialized degrader design.
| Selection Factor | Key Question | Practical Direction |
| E3 Ligase Type | Which E3 ligase system fits the research model and target context? | Compare CRBN, VHL, IAP, MDM2, or emerging ligase options based on project needs |
| Binding Motif | Does the ligand preserve key recognition contacts after modification? | Choose scaffolds with known modification tolerance or evaluate analogs carefully |
| Exit Vector | Can the linker be attached without disrupting E3 ligase engagement? | Prioritize attachment sites that point toward productive degrader geometry |
| Linker Compatibility | Does the ligand work with the required linker length and chemistry? | Assess PEG, alkyl, rigid, and mixed linker options when needed |
| Synthetic Route | Can ligand analogs and conjugates be prepared efficiently for comparison? | Use functionalized ligand building blocks or custom synthesis support |
| Optimization Stage | Is the project exploratory or focused on a defined degrader series? | Use broader ligand diversity early and matched analog sets later |
Selecting Ligands Based on E3 Ligase Type
The first decision is which E3 ligase system to recruit. CRBN and VHL ligands are often used as starting points because they support many established degrader design strategies. Other ligase systems may be valuable when researchers want to explore different protein contexts or alternative recruitment geometry. A useful overview of E3 ligand selection logic can be found in CRBN vs VHL: Choosing the Right E3 Ligase Ligand for Your PROTAC Project.
Evaluating Linker Attachment Sites and Exit Vectors
Linker attachment should preserve the E3 ligand recognition motif while presenting the linker toward a productive region of space. Exit vector choice can determine whether the target ligand and E3 ligase ligand are positioned favorably in the final degrader. If a ligand has multiple possible modification sites, a matched analog set can help clarify which exit vector supports better ternary complex formation.
Balancing Binding Affinity, Selectivity, and Synthetic Compatibility
Binding affinity is important, but it should not be the only selection criterion. A ligand with strong binding may still be difficult to use if it lacks a suitable attachment point or creates unfavorable degrader properties. Selectivity, synthetic compatibility, linker tolerance, and assay behavior should be evaluated together so that the selected ligand supports both chemical construction and functional interpretation.
Choosing Ligands for Early Screening or Focused Optimization
Early screening benefits from ligand diversity because the optimal E3 recruitment strategy may not be obvious. Focused optimization benefits from more controlled analog design, where ligand scaffold, linker attachment, and target ligand are changed systematically. BOC Sciences can support both approaches with catalog ligands, functionalized intermediates, and custom synthesis planning for specialized degrader projects.
Searching for High-Quality Ligands for E3 Ligases?
BOC Sciences provides a broad and reliable selection of in-stock ligands for E3 ligases to support PROTAC development and targeted protein degradation research. If you cannot find the ligand you need or require custom synthesis, our experts are here to help.
Request a Quote
Applications of E3 Ligase Ligands in Research
E3 ligase ligands are core building blocks for targeted protein degradation research. They enable researchers to construct bifunctional degraders, compare ligase recruitment strategies, evaluate target protein degradation, and develop chemical biology tools. Their value is greatest when the selected ligand can be connected to a coherent workflow that includes synthesis, assay design, and mechanism-focused interpretation.
PROTAC Discovery and Degrader Library Construction
E3 ligase ligands support PROTAC discovery by providing the recruitment element required for degrader assembly. In library construction, researchers may combine one or more E3 ligase ligands with different target ligands and linkers to explore chemical space. Related PROTAC library support can help organize degrader collections for screening and SAR development.
E3 Ligase Recruitment Strategy Comparison
Comparing E3 recruitment strategies allows researchers to test whether a target protein responds better to one ligase system than another. This comparison can clarify whether limited degradation arises from target ligand binding, linker geometry, E3 ligase choice, or protein context. E3 ligand comparison is especially useful when early degrader results are weak or inconsistent.
Target Protein Validation and Functional Studies
E3 ligase ligands can help build degraders used for target protein validation and functional studies. By reducing the abundance of a target protein in a controlled research setting, degrader tools can provide insights that complement genetic or inhibition-based approaches. When selecting ligands for this purpose, researchers should prioritize interpretable structure–activity trends and appropriate control compounds.
Chemical Biology Tools for TPD Platform Development
Ligands for E3 ligase are also valuable in chemical biology tool development. They may be used to prepare degrader probes, comparison molecules, homo-bifunctional ligase tools, or modular conjugates for mechanism-focused studies. For teams studying E3 systems directly, BOC Sciences provides related ubiquitin ligases products that can support protein degradation research workflows.
Ligand for E3 Ligase Product Options
BOC Sciences offers Ligand for E3 Ligase products to support PROTAC synthesis, degrader library construction, E3 recruitment comparison, and customized ligand development. Product options may include ready-to-use ligands, ligands bearing functional handles, ligand–linker intermediates, and custom structures designed around specific research objectives. These options help researchers reduce early design uncertainty and build a more coherent degrader development workflow.
Ready-to-Use E3 Ligase Ligands
Ready-to-use E3 ligase ligands are suitable for teams that have already selected a ligase system and need practical building blocks for degrader synthesis. These products can support early compound design, comparison of recruitment motifs, and analog preparation. BOC Sciences provides product information that helps researchers evaluate ligand class, structural features, and compatibility with downstream conjugation planning.
Ligands with Reactive Groups for PROTAC Synthesis
Functionalized E3 ligase ligands contain reactive groups that enable attachment to linkers or target ligand fragments. These products can simplify synthesis when the desired conjugation chemistry is already known. Reactive groups should be selected based on ligand scaffold tolerance, linker design, and the available coupling route. For additional chemistry support, BOC Sciences offers small molecule E3 ligase ligand services.
E3 Ligase Ligands for Linker Conjugation
E3 ligase ligands for linker conjugation help researchers prepare bifunctional degraders more efficiently. In some workflows, the E3 ligand is first connected to a linker and later coupled with the target-binding ligand. In other workflows, a preassembled target ligand–linker fragment is connected to the E3 ligand. BOC Sciences provides related E3 ligase ligand-linker conjugate products for modular degrader construction.
Custom Ligand Support for Specialized Degrader Designs
Some projects require E3 ligase ligands with specialized attachment sites, modified scaffolds, or matched analog series. Custom support can help researchers explore scaffold changes, linker-compatible handles, or less common recruitment strategies. BOC Sciences can discuss custom synthesis needs when catalog products do not fully match the design requirements of a degrader program.
Why Choose BOC Sciences Ligand for E3 Ligase Products?
BOC Sciences supports E3 ligase ligand research with product options and technical capabilities aligned with degrader design workflows. Our focus is to help customers choose ligands that are chemically practical, mechanistically relevant, and suitable for their intended research stage. From catalog selection to custom ligand planning, BOC Sciences provides support for teams developing PROTACs and related targeted protein degradation tools.
 Broad Selection for Diverse E3 Ligase Recruitment Strategies
Broad Selection for Diverse E3 Ligase Recruitment Strategies
BOC Sciences provides E3 ligase ligand options covering commonly used recruitment systems and exploratory ligand classes. This enables researchers to compare different ligase strategies and select building blocks that fit their degrader design goals.
 Technical Support for Ligand Selection and Project Planning
Technical Support for Ligand Selection and Project Planning
Our team can discuss ligand class, linker attachment, exit vector selection, conjugation route, and project stage. This helps researchers narrow the design space and plan informative analog sets for screening or optimization.
 Custom Synthesis Support for Specialized Ligand Requirements
Custom Synthesis Support for Specialized Ligand Requirements
When catalog products do not meet project needs, BOC Sciences can support custom synthesis of modified E3 ligase ligands, functionalized intermediates, or ligand analogs designed for specific degrader construction routes.
 Research-Oriented Product Information for Efficient Procurement
Research-Oriented Product Information for Efficient Procurement
Clear product information helps scientists and procurement teams evaluate ligand class, functional handles, intended research use, and compatibility with downstream synthesis. This supports smoother communication between scientific planning and purchasing decisions.
Frequently Asked Questions (FAQ)

Still have questions?
Contact Us
How do I choose the right E3 ligase ligand?
Selecting an appropriate E3 ligase ligand depends on your target protein, degradation strategy, and experimental system. Researchers typically consider ligand compatibility with known E3 ligases such as CRBN or VHL, linker attachment feasibility, and structural data availability. It is also important to evaluate physicochemical properties that influence cell permeability and stability. Many procurement teams prioritize suppliers that provide detailed characterization data, scalable synthesis capability, and technical consultation. BOC Sciences supports ligand selection by offering a diverse portfolio and guidance aligned with targeted protein degradation workflows.
What are common applications of E3 ligase ligands in research?
E3 ligase ligands are widely used in targeted protein degradation studies, particularly in PROTAC and molecular glue development. They serve as essential components that recruit E3 ubiquitin ligases to induce selective degradation of disease-related proteins. Beyond oncology-focused research, they are also applied in neuroscience, immunology, and epigenetics studies. Researchers use these ligands to validate targets, study protein function, and explore degradation pathways. Reliable sourcing and consistent quality are critical for reproducibility, making supplier expertise an important factor in procurement decisions.
Can E3 ligase ligands be customized for specific targets?
Yes, customization is a common requirement in advanced research projects involving targeted degradation. Scientists often need modified ligands with specific functional groups for linker conjugation or improved binding characteristics. Custom synthesis providers can optimize molecular structures based on project requirements, including scaffold modification and structure-activity relationship exploration. BOC Sciences offers flexible customization services, supporting researchers with tailored ligand design and synthesis to align with specific experimental objectives, helping streamline early-stage discovery workflows without compromising adaptability.
What factors impact performance of E3 ligase ligands?
The performance of an E3 ligase ligand is influenced by multiple factors including binding affinity to the ligase, chemical stability, solubility, and compatibility with linker chemistry. Structural orientation plays a key role in forming an effective ternary complex when used in PROTAC systems. Additionally, cellular context such as E3 ligase expression levels can affect degradation efficiency. Researchers often conduct iterative optimization to balance these variables. Access to well-characterized ligands and technical insights from experienced suppliers can significantly improve experimental outcomes.
What should buyers consider when sourcing E3 ligase ligands?
Procurement teams typically evaluate supplier reliability, product diversity, technical support, and scalability when sourcing E3 ligase ligands. Availability of documentation, analytical data, and customization options are also critical considerations. For research organizations, aligning supply capabilities with long-term project needs is essential. BOC Sciences provides a comprehensive range of ligands along with technical support to assist decision-making, helping research teams ensure continuity and consistency in their targeted protein degradation studies while maintaining operational efficiency.
Client Feedback on Ligand for E3 Ligase Products
Helpful Selection for E3 Recruitment Studies
“The E3 ligase ligand options from BOC Sciences helped our chemistry team compare recruitment strategies across several degrader designs. The product information was clear enough for both scientific review and procurement planning.”
— Discovery Chemistry Lead, North America
Useful Discussion Around Linker Attachment
“We needed to evaluate whether a selected E3 ligand could tolerate linker installation at a specific position. BOC Sciences provided practical technical communication that supported our synthesis planning and analog prioritization.”
— Senior Scientist, European Research Group
Support for Custom Ligand Requirements
“Our project required modified E3 ligase ligand structures for a focused degrader series. The ability to discuss custom synthesis needs with BOC Sciences helped us align compound design with our research workflow.”
— Project Manager, Life Science R&D Team
Efficient Procurement Communication
“The product category was well organized, and the technical details helped our team distinguish ligand-only products from ligand-linker conjugates. This reduced internal clarification and made ordering discussions more efficient.”
— Procurement Specialist, Research Organization
Discover More Research Products
Explore featured products that can expand your research options and accelerate your next discovery.
Expert Services to Move Your Project Forward
Access end-to-end service solutions that help bring efficiency, flexibility, and expertise to your research pipeline.
Insights and Resources
News
Technical Information











![3-(1-oxo-3,5,6,7-tetrahydropyrrolo[3,4-f]isoindol-2-yl)piperidine-2,6-dione](https://resource.bocsci.com/structure/2616539-03-6.gif)









 Fig.1 SNIPER-Mediated IAP-Dependent Target Protein Degradation Workflow (BOC Sciences Original).
Fig.1 SNIPER-Mediated IAP-Dependent Target Protein Degradation Workflow (BOC Sciences Original).