What Are Molecular Glues?
Molecular glues are a class of small-molecule compounds that chemically induce or stabilize protein–protein interactions (PPIs) that would not otherwise occur under normal physiological conditions. By bridging two proteins at a shared interface, a molecular glue creates a stable ternary complex that redirects cellular machinery toward a specific biochemical outcome—most notably the ubiquitin-mediated degradation of a target protein. First conceptualized through observations with natural products such as cyclosporin A, FK506, and rapamycin, the molecular glue concept was later validated in oncology with the discovery that immunomodulatory drugs (IMiDs)—thalidomide, lenalidomide, and pomalidomide—act by recruiting neosubstrates to the E3 ubiquitin ligase cereblon (CRBN). Today, rationally designed molecular glue degraders (MGDs) represent one of the most rapidly advancing modalities in targeted protein degradation (TPD) research.
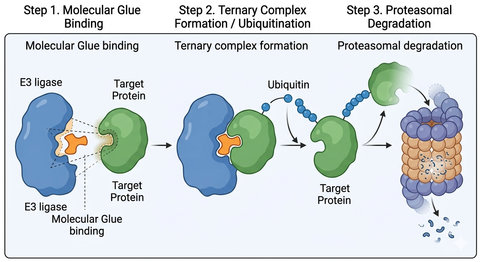 Fig.1 Molecular Glue Induced Targeted Protein Degradation (BOC Sciences Original).
Fig.1 Molecular Glue Induced Targeted Protein Degradation (BOC Sciences Original).
Physicochemical Properties of Molecular Glues
Molecular glues stand apart from other targeted protein degraders primarily through their compact, drug-like physicochemical profiles:
- Low molecular weight: Typically < 500 Da, fully compliant with Lipinski's Rule of Five, enabling favorable absorption and distribution properties in cell-based assays.
- High ligand efficiency: A single small molecule simultaneously engages two protein surfaces, achieving remarkable biological potency per unit molecular mass.
- Monovalent architecture: No linker or bifunctional scaffold is required; the compound itself serves as the entire pharmacophore, simplifying synthetic optimization.
- Favorable solubility and permeability: Compact size supports membrane permeability across a wide range of in vitro model systems, facilitating cellular uptake in research contexts.
- Sub-stoichiometric activity: Because the compound is released after triggering ubiquitination and is not consumed in the degradation cycle, a single molecule can catalytically mediate multiple rounds of target protein degradation—providing potent activity at low concentrations.
- Conformational plasticity: Molecular glues often engage a cryptic or shallow binding groove that becomes fully ordered only upon ternary complex formation, a phenomenon sometimes called "induced fit" interface stabilization.
Mechanism of Action of Molecular Glue Degraders
The canonical mechanism of a molecular glue degrader (MGD) proceeds through four sequential steps:
Step 1 Ligase Binding
The molecular glue binds to an E3 ubiquitin ligase substrate receptor (e.g., CRBN, DCAF15, or DDB1), typically occupying a shallow hydrophobic groove with modest intrinsic affinity.
Step 2 Neosubstrate Recruitment
Binding of the glue molecule reshapes the ligase surface, generating a composite interface that is complementary to a specific degron motif on the target (neosubstrate) protein. The neosubstrate is then recruited into a stable ternary complex: [E3 ligase – molecular glue – neosubstrate].
Step 3 Ubiquitin Transfer
Within the ternary complex, E2 ubiquitin-conjugating enzymes transfer ubiquitin chains onto accessible lysine residues on the neosubstrate. Productive polyubiquitination requires sufficient conformational accessibility of the target's lysine residues relative to the E2 active site.
Step 4 Proteasomal Degradation and Glue Recycling
The polyubiquitinated neosubstrate is recognized by the 26S proteasome, unfolded, and degraded into short peptides. The molecular glue is then released and recycled, enabling additional rounds of target recruitment and degradation—conferring catalytic-like efficiency.
Structural Features of Molecular Glues
Molecular glues are low-molecular-weight, structurally compact small molecules, typically featuring mono- or bicyclic scaffolds with rigid cores. Their core structure contains two key pharmacophoric regions: one end serves as the E3 ligase recognition module (such as the glutarimide ring in CRBN ligands), while the other interacts with the target protein interface. These are connected by short alkyl or aryl linkers, enabling precise control of protein proximity and triggering conformational changes in the target protein, effectively “gluing” together proteins that would not otherwise interact. Unlike the bifunctional, large-molecule design of PROTACs, molecular glues eliminate long flexible linkers and adopt a more globular, low-flexibility conformation, offering improved oral bioavailability and drug-like properties.
Structure-Function Relationship of Molecular Glues
E3 Ligase Recognition Module (Anchoring Function)
The E3 ligase recognition module forms the structural basis of a molecular glue. Its main role is to anchor the molecule to a specific E3 ubiquitin ligase and position it for target protein recruitment. A typical example is the glutarimide ring, which fits into the hydrophobic pocket of CRBN and is stabilized by hydrogen bonds with residues such as Trp403 and His379. In DCAF15-directed molecular glues, sulfonamide scaffolds bind the WD40 domain through hydrophobic and hydrogen-bonding interactions. Overall, this module acts as a positioning anchor that secures the glue on the E3 ligase surface while preserving an interface for target binding.
Target Protein Interface Adaptation Module (Recruitment Function)
The target protein interface adaptation module determines substrate selectivity and functional outcome. This region is often flexible and contains substituents such as aryl, amino, or alkyl groups that extend toward the target protein surface to create new hydrophobic or polar interactions. These contacts can trigger conformational changes and promote formation of a new binding interface. For example, C5 aryl extensions may interact with the GTPase domain of GSPT1, while C4 amino substituents can complement the hydrophobic surface of zinc finger proteins such as IKZF1/3. In some cases, spirocyclic or bridged ring systems improve three-dimensional complementarity and cooperativity. Overall, this module recruits target proteins through an induced-fit mechanism.
Bridge Module (Spacing Control Function)
The bridge module links the E3 ligase-binding end and the target-adapting end, controlling the distance and orientation between the two proteins. This is essential for efficient ternary-complex formation. Common bridge elements include short alkyl chains, aromatic rings, and heteroaromatic rings. Short alkyl chains can adjust the Cα distance between ligase and target, while aromatic systems increase rigidity, reduce entropic loss, and may add pi-stacking interactions. Oxygen or nitrogen atoms can further tune polarity and solubility, and stereocenters may strongly affect chiral fit in the ternary complex. In this way, the bridge acts as a molecular ruler that influences ubiquitination efficiency.
Specific Structure-Function Relationships
The activity of molecular glues depends on highly precise structure–function relationships, and even small structural changes can strongly affect substrate selectivity and degradation profile. In CRBN-based molecular glues, changes at the C5 position can switch activity from IKZF1/3 degradation to selective GSPT1 degradation, emphasizing the importance of interface size and hydrophobic complementarity. The rigid cyclic structure of the glutarimide ring is also crucial for CRBN binding, and disrupting this rigidity can eliminate activity. In DCAF15-directed molecular glues, the sulfonamide motif is essential for proper molecular orientation through hydrogen bonding, while aryl substitution patterns influence selectivity toward substrates such as RBM39. Overall, molecular glue function arises from precise geometric matching and interaction remodeling at the protein–protein interface.
Classification of Molecular Glues
Molecular glues can be broadly categorized by their E3 ligase recruitment mechanism, structural scaffold, or mode of PPI modulation:
IMiDs and CELMoDs (CRBN-based)
This is the most extensively studied class of molecular glues. IMiDs such as thalidomide, lenalidomide, and pomalidomide, along with next-generation CELMoDs including iberdomide, eragidomide, and mezigdomide, share the glutarimide pharmacophore and function through CRBN. These compounds recruit neosubstrates such as IKZF1/3, CK1α, GSPT1, RBM39, and cyclin K to the CRL4 CRBN E3 ligase complex.
Sulfonamide-Based MGDs (DCAF15-based)
Sulfonamide-based molecular glue degraders represent a well-defined DCAF15-dependent class. Compounds such as indisulam, E7820, and tasisulam bind the WD40 receptor DCAF15 within the CRL4 DCAF15 ligase complex and promote selective degradation of RBM39. Their activity relies on induced ternary-complex formation and recognition of the RRM2 degron motif within the recruited splicing factor.
Next-Generation Rational MGDs
Next-generation rational MGDs are being discovered through DEL screening, ASMS, and structure-based design. Compared with classical molecular glues, they expand induced proximity to additional ligase systems, including DCAF16, DDB1, and VHL-associated complexes. Representative examples include (R)-CR8, optimized indisulam analogs, and newer programs targeting cyclin K, CDK12, NEK7, BRD9, VAV1, RAF1, and KRAS.
Natural-Product-Derived Molecular Glues (Stabilizers)
A distinct subclass of molecular glues is derived from natural products and primarily stabilizes native protein–protein interactions rather than recruiting neosubstrates for degradation. Classic examples include rapamycin, which bridges FKBP12 and mTOR, and FK506, which stabilizes FKBP12–calcineurin. In plant systems, auxin enables TIR1-dependent recruitment of AUX/IAA proteins and supports auxin-inducible degron technology.
Key Molecular Glue Targets and E3 Ligase Systems

Common Targets
Molecular glues primarily act on proteins that have traditionally been considered difficult to target, as these proteins often lack a defined active site or deep hydrophobic pocket and are commonly transcription factors, splicing factors, or signaling regulators. By inducing new interaction interfaces between these proteins and E3 ligases, molecular glues enable selective degradation or functional modulation and are widely used in mechanistic research. Representative targets include:
- IKZF1 / IKZF3
- GSPT1
- RBM39
- CK1α
- Cyclin K / CDK12
- BRD4
- VAV1
- RAF1

Common E3 Ligases
E3 ligases are the core effector systems in molecular glue activity, responsible for recognizing and ubiquitinating target proteins and directing them to the proteasomal degradation pathway. Different E3 ligase systems show distinct structural features and substrate selectivity, making them a central focus in molecular glue design and research. Representative E3 ligases include:
- CRBN (Cereblon)
- DCAF15
- DCAF16
- DDB1 / CRL4 complex
- VHL (Von Hippel–Lindau)
- MDM2
- RNF4
- TRIM21
Molecular Glues vs. PROTACs
Both molecular glues and PROTACs (Proteolysis-Targeting Chimeras) exploit the ubiquitin–proteasome system to degrade target proteins, but they differ fundamentally in structure, mechanism, and practical utility for research applications.
| Parameter | Molecular Glues | PROTACs |
| Architecture | Monovalent; single small molecule | Bifunctional; two ligands connected by a linker |
| Molecular Weight | Typically < 500 Da | Typically 700–1,100 Da |
| Mechanism | Induces a de novo (neomorphic) PPI | Bridges independently existing binding pockets |
| Design Requirement | No pre-existing binder for target required | Requires known binder for both target and E3 ligase |
| Hook Effect | Absent or minimal | Pronounced at high concentrations |
| Ternary Complex Cooperativity | Emergent (glue creates the interface) | Dependent on linker geometry and length |
| Synthetic Complexity | Generally lower (single scaffold) | Higher (bifunctional, linker optimization required) |
| Physicochemical Profile | Favorable; Rule-of-Five compliant | Often beyond Rule-of-Five; permeability challenges |
| Target Scope | Proteins lacking known small-molecule binders | Proteins with tractable binding pockets |
| Discovery Strategy | Phenotypic screening, DEL, ASMS, serendipity | Structure-based rational design more tractable |
Research implication: For targets that lack a characterized small-molecule binding site, molecular glues offer a compelling alternative to PROTACs, since they do not require a pre-existing ligand for the protein of interest. Conversely, PROTACs benefit from a more structurally transparent design logic when both target and E3 ligase binders are already known.
Advantages of Molecular Glues in Research Applications
01Ability to Target Previously Intractable Proteins
Molecular glues can engage proteins that lack a catalytic site or deep hydrophobic pocket by inducing protein–protein interactions. This makes transcription factors, scaffolding proteins, and intrinsically disordered regions accessible research targets.
02Novel Mechanism of Action Through Induced Proximity
Unlike conventional inhibitors, molecular glues function by rewiring protein interaction networks. This mechanism is particularly valuable for studying selective protein degradation mediated by E3 ligases.
03Compact Structures with Broad Optimization Potential
As monovalent small molecules, molecular glues are typically structurally compact and do not require complex linker design. This makes them well suited for medicinal chemistry optimization, structure–activity relationship studies, and library screening.
04Potential for Highly Selective Degradation
Molecular glues rely on ternary-complex geometry and cooperativity, which can enable highly selective substrate degradation. In some cases, this selectivity goes beyond what would be predicted from binding affinity alone.
05Compatibility with Diverse Research Platforms
Molecular glues can be broadly applied in protein degradation studies, chemoproteomics, structural biology, and high-throughput screening. They support the full workflow from mechanistic investigation to functional validation.
06Expansion of E3 Ligase Research Space
Molecular glues provide useful tools for exploring substrate recognition and functional mechanisms across different E3 ligases. They also support the discovery of new E3–substrate relationships and the development of novel degradation strategies.
Applications of Molecular Glues
Cancer-Related Protein Degradation Research
Molecular glues are widely used to study the selective degradation of cancer-associated proteins, especially transcription factors, splicing factors, and signaling regulators that are difficult to modulate by conventional approaches.
Immune and Inflammation Pathway Research
In immune and inflammation research, molecular glues can be used to modulate the stability of critical regulatory proteins, including transcription factors and signaling adaptors. This makes them valuable for studying immune cell function, pathway regulation, and the molecular basis of inflammatory signaling.
Research Tools for TPD Platforms
Molecular glues are important tools in targeted protein degradation research. They can be used to evaluate E3 ligase function, compare different degradation strategies such as molecular glues and PROTACs, and support the development and optimization of induced-proximity degradation platforms.
Protein Function Validation and Knockdown
Molecular glues can serve as chemical knockdown tools by rapidly and selectively reducing the level of a target protein. This makes them highly useful for protein function validation, especially in systems where genetic manipulation is less efficient or where reversible protein depletion is preferred.
Why Choose BOC Sciences Molecular Glue Products?
 Broad Product Selection
Broad Product Selection
BOC Sciences offers molecular glue products covering major systems such as CRBN and DCAF15, as well as a range of emerging targets for different protein degradation research needs.
 Clear Technical Information
Clear Technical Information
Each product page includes key information such as target, mechanism, physicochemical properties, and storage conditions to help researchers evaluate and select suitable compounds more efficiently.
 Reliable and Consistent Quality
Reliable and Consistent Quality
Standardized quality control and consistent product quality provide dependable support for protein degradation studies and mechanism-based research.
 Support for Research Project Expansion
Support for Research Project Expansion
In addition to standard catalog products, BOC Sciences can provide flexible support for molecular glue-related research and project development.
Frequently Asked Questions (FAQ)

Still have questions?
Contact Us
What is a molecular glue and its role in drug development?
Molecular glue is a small molecule that induces or stabilizes interactions between two proteins, regulating biological processes. In drug development, it is mainly used in targeted protein degradation (TPD) to recruit E3 ubiquitin ligases for selective protein degradation. BOC Sciences provides high-purity molecular glues and custom synthesis services to accelerate early-stage drug discovery and mechanistic studies.
How does a molecular glue differ from PROTACs?
PROTACs are bifunctional molecules linking a target protein to an E3 ligase, whereas molecular glues are single small molecules that enhance or create protein-protein interactions for the same degradation effect. Molecular glues generally have smaller size and better pharmacokinetics. BOC Sciences offers both PROTAC and molecular glue supply and custom development services to support fast experimental access.
How are effective molecular glue candidates screened?
Screening typically involves protein-E3 binding assays (SPR, ITC), cellular degradation assays, and structural optimization. High-throughput screening (HTS) and computer-aided drug design (CADD) improve hit rates. BOC Sciences provides chemical libraries and custom synthesis to support candidate screening and structural optimization, reducing R&D risk.
What challenges do molecular glues face in clinical development?
Challenges include target specificity, pharmacokinetics, and biological complexity across cell types. BOC Sciences offers high-purity compounds, custom synthesis, and early ADME/Tox support to help evaluate candidate feasibility.
Which disease areas are suitable for molecular glue drug development?
Molecular glues show potential in cancer, autoimmune diseases, and neurodegenerative disorders by selectively degrading disease-relevant proteins. BOC Sciences supports clients with compound supply and custom development services across research to preclinical stages.
Client Testimonials for Our Molecular Glue Products
Cost-Effective Product Offering
“BOC Sciences provides molecular glue products at very competitive prices without compromising quality, which helped us efficiently manage our early discovery budget.”
— Dr. Anderson, Senior Scientist at a US-based Biotech
Reliable Product Quality
“The molecular glue compounds we received from BOC Sciences showed excellent purity, stability, and batch-to-batch consistency, fully meeting the standards required for our research program.”
— Ms. Keller, Principal Investigator at a European Research Institute
Broad Specification Selection
“BOC Sciences offers a wide range of molecular glue products with diverse specifications, making it easy for us to identify suitable candidates for different experimental needs.”
— Dr. Morgan, Project Leader at a UK-based Pharmaceutical Company
Flexible Custom Synthesis Support
“We were impressed by the custom synthesis options provided by BOC Sciences, which allowed us to obtain tailored molecular glue compounds aligned with our project requirements.”
— Mr. Tanaka, R&D Manager at a Japan-based Biotech Firm
Discover More Research Products
Explore featured products that can expand your research options and accelerate your next discovery.
Expert Services to Move Your Project Forward
Access end-to-end service solutions that help bring efficiency, flexibility, and expertise to your research pipeline.
News
Technical Information





















 Fig.1 Molecular Glue Induced Targeted Protein Degradation (BOC Sciences Original).
Fig.1 Molecular Glue Induced Targeted Protein Degradation (BOC Sciences Original).