Project Background
A biotechnology client had synthesized a focused series of BRD4-targeting PROTAC molecules derived from a common bromodomain warhead but with different linker lengths, linker attachment positions, and CRBN ligand variants. Although several compounds showed similar biochemical affinity, the client could not determine which molecules were most likely to deliver robust intracellular degradation and requested a structured in vitro ranking workflow.
Technical Challenges
The main difficulty was that affinity data alone did not correlate with cellular performance. Some compounds showed acceptable binding but weak degradation, while others displayed non-linear dose response suggestive of hook effect behavior. The client needed clarity on whether the limiting factors were ternary complex productivity, permeability, or degradation kinetics.
BOC Sciences Solutions
- Stepwise assay cascade: We established a tiered workflow beginning with binary binding confirmation, followed by ternary complex assessment, then cell-based degradation studies and property-supporting analysis.
- Comparative degradation profiling: Twelve degraders were evaluated across concentration and time-course studies to measure DC50, Dmax, onset rate, and recovery behavior.
- Root-cause analysis: Orthogonal results were integrated to determine which candidates were limited by weak complex cooperativity and which were limited by cellular exposure.
Project Outcomes
The client received a clear rank order for all 12 molecules, with 3 compounds identified as strong leads. The top degrader delivered submicromolar DC50 and greater than 85% BRD4 degradation in the primary screening cell line, while two other compounds were deprioritized despite similar affinity because they formed less productive ternary complexes. A separate subgroup with acceptable complex formation but poor cellular output was flagged for chemistry revision focused on exposure-related properties. This study reduced optimization ambiguity and gave the client a mechanism-backed path forward.
Project Background
A pharmaceutical partner was developing a PROTAC program against an EGFR-pathway signaling node and had observed inconsistent protein-loss data across different cell models. The team needed to confirm whether the apparent degradation phenotype was genuine, whether it was dependent on recruited ligase biology, and which assay format would best support future screening of analogs.
Technical Challenges
Preliminary Western blot results showed partial target reduction, but the magnitude varied by cell line and dosing window. The client also lacked clarity on whether poor reproducibility reflected assay variability, target turnover differences, or insufficient intracellular engagement.
BOC Sciences Solutions
- Model selection strategy: We profiled multiple cell backgrounds with different target and ligase expression features to identify the most informative evaluation system.
- Mechanism confirmation package: The project combined target engagement-supporting readouts, ternary complex studies, ubiquitination analysis, and proteasome-dependency controls.
- Kinetic optimization: Several treatment windows and concentrations were compared to define the most discriminating conditions for lead ranking and reproducibility.
Project Outcomes
Our workflow showed that the client's lead series produced meaningful degradation only in cellular contexts with sufficient ligase support and adequate intracellular exposure. We identified one cell model as the optimal screening background and established assay conditions that improved signal consistency substantially. Across 18 tested compounds, 4 molecules showed strong mechanism-consistent degradation and 1 candidate emerged as the preferred lead because it combined deeper target loss, cleaner selectivity behavior, and more sustained degradation over time. The client used these findings to refine both assay strategy and analog design priorities.

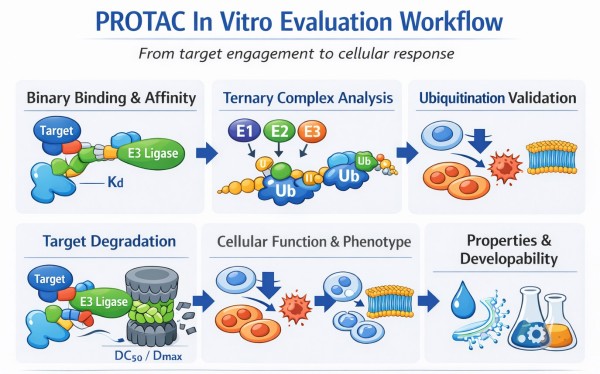 Fig.1 PROTAC in vitro evaluation workflow diagram (BOC Sciences).
Fig.1 PROTAC in vitro evaluation workflow diagram (BOC Sciences).