What Are PROTACs?
PROTACs, or proteolysis targeting chimeras, are bifunctional small-molecule degraders designed to bring a protein of interest into proximity with an E3 ubiquitin ligase. Instead of simply blocking the activity of a protein, a PROTAC is intended to promote selective removal of that protein through the ubiquitin-proteasome system. This degradation-centered mechanism has made PROTAC technology an important part of targeted protein degradation research, especially for teams seeking chemical tools that can reduce the abundance of proteins involved in signaling, transcriptional control, epigenetic control, and other complex biological processes.
For research groups working in drug discovery, chemical biology, and protein function studies, PROTACs provide a way to evaluate what happens when a target protein is depleted rather than inhibited. BOC Sciences provides PROTAC-related products, building blocks, and technical resources that help researchers explore degrader design, linker selection, target engagement, E3 ligase recruitment, and degradation assessment across diverse experimental workflows.
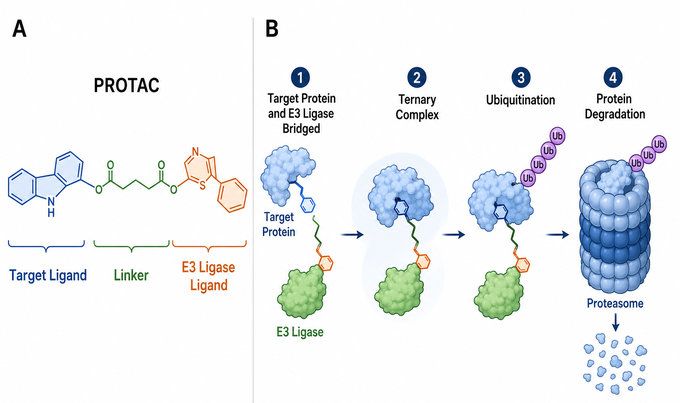 Fig.1 PROTAC-mediated targeted protein degradation workflow (BOC Sciences Original).
Fig.1 PROTAC-mediated targeted protein degradation workflow (BOC Sciences Original).
Definition of Proteolysis Targeting Chimeras
A PROTAC is typically composed of three functional elements: a ligand that binds the protein of interest, a ligand that recruits an E3 ubiquitin ligase, and a chemical linker connecting the two ligands. This modular architecture enables the compound to form a ternary complex containing the target protein, the PROTAC molecule, and the recruited E3 ligase. Once the complex is positioned in a productive geometry, the target protein can be ubiquitinated and directed to proteasome-mediated degradation.
The term proteolysis targeting chimera reflects this hybrid design. One end of the molecule recognizes the target protein, another end engages the ubiquitination machinery, and the linker controls distance, orientation, flexibility, and physicochemical properties. Because these three components influence each other, PROTAC design requires careful balancing of binding affinity, ternary-complex cooperativity, permeability, solubility, linker geometry, and cellular degradation response.
Key Differences Between PROTAC Degraders and Conventional Inhibitors
Conventional small-molecule inhibitors generally act through occupancy-driven pharmacology. They bind to an active site, allosteric pocket, or protein interface and reduce a specific biochemical activity while the compound remains associated with the target. PROTAC degraders follow a different logic. They are event-driven molecules that bring the target protein close to an E3 ligase, trigger ubiquitination, and allow the cell's proteasomal machinery to remove the target. After the degradation event, the PROTAC may participate in additional rounds of target recruitment.
This distinction affects how researchers evaluate a PROTAC. Binding affinity alone is not sufficient to predict degradation performance. Productive ternary-complex formation, E3 ligase compatibility, cellular localization, linker geometry, target lysine accessibility, and proteasome dependence all contribute to the final degradation profile. For this reason, PROTAC optimization often combines medicinal chemistry, protein biochemistry, cellular assays, and structural analysis rather than relying on a single binding measurement.
Mechanism of PROTAC-Mediated Protein Degradation
The PROTAC mechanism is an induced-proximity process that redirects the ubiquitin-proteasome system toward a selected protein of interest. The process generally includes ternary complex formation, target ubiquitination, proteasome recognition, and final proteolysis.
Step 1: Ternary Complex Formation
A PROTAC molecule simultaneously binds the protein of interest and an E3 ubiquitin ligase to form a target protein-PROTAC-E3 ternary complex. This step is often critical for degradation efficiency, as the stability, binding affinity, cooperativity, and conformation of the ternary complex determine whether the recruited proteins are positioned productively for downstream ubiquitination.
Step 2: Target Protein Ubiquitination
After ternary complex formation, the recruited E3 ligase works with an E2 ubiquitin-conjugating enzyme to transfer ubiquitin molecules to lysine residues on the target protein through isopeptide bonds. Repeated transfer can generate a polyubiquitin chain, commonly K48-linked, which serves as a degradation signal for the target protein.
Step 3: Proteasome Recognition
The polyubiquitinated target protein is recognized by the 26S proteasome, mainly through the 19S regulatory particle. The ubiquitin chain functions as a degradation tag, helping direct the marked target protein toward the proteasomal core for processing.
Step 4: Proteolysis and Ubiquitin Recycling
Once recognized, the target protein is unfolded and translocated into the 20S core particle, where catalytic threonine protease sites cleave it into peptide fragments. This ATP-dependent process is accompanied by ubiquitin removal and recycling through deubiquitinating enzymes.
Core Structural Components of PROTACs
PROTAC molecules are modular, but modularity does not mean that each component can be optimized independently. The target ligand, E3 ligase ligand, linker, and conjugation chemistry collectively determine the three-dimensional shape, flexibility, cellular exposure, and degradation outcome. Even small changes in linker length or attachment position can alter ternary-complex geometry and switch a compound from active to inactive.
Ligands for Target Protein Engagement
The target ligand anchors the PROTAC to the protein of interest. Useful ligands are typically selected based on binding mode, availability of a modification site, selectivity profile, and compatibility with conjugation. A ligand with high affinity is often useful, but it is not always sufficient. The ligand must tolerate linker installation at a position that does not disrupt target binding and allows the E3 ligase to approach the target in a productive orientation.
BOC Sciences offers ligands for target protein research and related resources for teams assembling degrader candidates from known target-binding chemotypes.
E3 Ligase Ligands for Degrader Construction
The E3 ligase ligand determines which ubiquitin ligase is recruited by the PROTAC. Common ligase-binding motifs can vary in size, polarity, stereochemical requirements, and tolerance for linker attachment. These characteristics directly affect the design of the full degrader. For example, a ligand with a well-defined exit vector may simplify conjugation, while a ligand with stricter spatial requirements may require more extensive linker exploration.
Because E3 ligase expression and localization can differ by experimental system, researchers often compare more than one ligase-recruiting strategy for a target of interest. Productive degradation is therefore a function of both chemical design and biological context.
PROTAC Linkers and Linker Optimization
The linker is not a passive spacer. It controls the distance between the two ligands, the conformational freedom of the molecule, the orientation of recruited proteins, and many physicochemical properties relevant to cellular studies. Common linker chemistries include alkyl chains, polyethylene glycol motifs, piperazine-containing linkers, triazoles, amides, and other heteroatom-containing structures used to tune polarity and flexibility.
Linker optimization is often one of the most important stages of PROTAC development. A shorter linker may force a compact ternary complex, while a longer linker may improve reach but increase flexibility and reduce productive complex formation. BOC Sciences provides PROTAC linker products and linker design and optimization services to support systematic exploration of linker effects.
Functional Groups for Conjugation and Library Design
Functional groups used for conjugation influence synthetic feasibility, chemical stability, and the speed of degrader library preparation. Amine, carboxyl, alkyne, azide, hydroxyl, halide, and activated ester handles are commonly considered when designing conjugatable building blocks. The choice of chemistry should preserve ligand recognition while allowing efficient assembly of a structurally diverse PROTAC series.
For library design, researchers often vary linker length, polarity, rigidity, and attachment positions in parallel. A carefully planned library can reveal structure-activity relationships that would be missed by testing only one or two molecules. BOC Sciences supports these workflows through PROTAC library resources and related custom synthesis capabilities.
Common E3 Ligase Systems Used in PROTAC Design
E3 ligase selection is a central decision in PROTAC design. The recruited ligase must be compatible with the target protein, expressed in the experimental system, and able to position ubiquitination machinery near accessible regions of the target. Different E3 ligase systems present different advantages for degrader research, and comparing several ligase recruitment approaches can be valuable when the first design series does not produce sufficient target degradation.
Cereblon-Based PROTACs
Cereblon-based PROTACs are widely used in targeted protein degradation research because cereblon ligands are well established as recruiter motifs and are compatible with many bifunctional degrader designs. Cereblon-recruiting PROTACs often use glutarimide-containing ligands such as thalidomide-derived or related structures as the E3-binding component. The cereblon ligand is typically connected to a target binder through a linker installed at a position that preserves ligase recognition.
Designing a cereblon-based PROTAC requires attention to ligand attachment, ternary-complex geometry, target protein accessibility, and potential effects of linker bulk on cellular behavior. BOC Sciences provides resources for CRBN-based PROTAC development to help researchers evaluate this commonly used ligase strategy.
VHL-Based PROTACs
VHL-based PROTACs are another major class of degraders. VHL ligands are frequently used when researchers want to compare degradation profiles across ligase systems or when the target protein may be better positioned for ubiquitination through a VHL-recruiting architecture. VHL ligands often have distinct stereochemical and polarity characteristics compared with cereblon ligands, which can affect linker selection and full-molecule properties.
VHL recruitment can be particularly useful for systematic studies of ternary-complex cooperativity because structural modeling and biophysical analysis can guide ligand orientation and linker design. BOC Sciences supports VHL-based PROTAC development with design and synthesis capabilities aligned with research workflows.
IAP- and MDM2-Recruiting PROTACs
IAP- and MDM2-recruiting PROTACs broaden the range of E3 ligase systems available for induced protein degradation studies. IAP-recruiting degraders are often discussed in relation to inhibitor of apoptosis protein ligase recruitment, while MDM2-recruiting approaches provide another framework for exploring target degradation through alternative ligase engagement. These systems can offer different cellular localization, expression patterns, and ubiquitination geometries.
For some targets, comparing cereblon, VHL, IAP, and MDM2 recruiters may help researchers understand whether a lack of degradation reflects target biology, linker geometry, or ligase mismatch. BOC Sciences provides support for IAP-based PROTAC development and MDM2-based PROTAC development as part of broader degradation platform research.
Emerging E3 Ligase Ligands for Expanded Degrader Space
Although several E3 ligases dominate current PROTAC research, the human ubiquitin ligase system contains many potential recruitment opportunities. Emerging E3 ligase ligands may expand the target scope of PROTAC technology, improve compatibility with specific cell systems, or enable degradation of proteins that are not efficiently addressed by more established recruiters.
Exploration of emerging ligases requires ligand discovery, ligase expression analysis, structural evaluation, and assay development. Researchers may use small-molecule screening, covalent ligand discovery, proteomics, and computational modeling to identify new recruiter motifs. BOC Sciences offers design of the ligase system support and related ligand resources for teams investigating new E3 recruitment strategies.
PROTAC Design Strategies for Drug Discovery Research
PROTAC design is an iterative process that integrates target biology, ligand chemistry, linker engineering, ternary-complex analysis, and degradation evaluation. A successful design workflow usually begins with a clear target hypothesis and a practical assessment of available binders. It then proceeds through modular synthesis, cellular screening, mechanistic confirmation, and structure-activity relationship analysis.
01Target Selection and Ligand Availability
Target selection should consider the biological question, protein abundance, subcellular localization, turnover rate, available binding pockets, and the presence of accessible lysine residues. PROTACs are especially useful when researchers want to evaluate target removal rather than simple activity modulation. However, a practical PROTAC project still requires a ligand that can engage the target and tolerate conjugation.
When a high-quality ligand is not available, researchers may begin with fragment screening, virtual screening, protein-ligand modeling, or custom ligand design. BOC Sciences provides ligand design for target protein support to help teams identify or refine target-binding starting points for degrader construction.
02Linker Length, Composition, and Exit Vector Design
The linker must connect the two ligands while maintaining productive protein-protein proximity. Length determines the reach between the target protein and E3 ligase, while composition influences flexibility, solubility, permeability, and conformational preference. Exit vectors determine the direction in which each ligand projects the linker, shaping the architecture of the ternary complex.
A good linker design strategy usually includes a focused set of lengths and chemistries rather than random variation. Researchers may combine alkyl, PEG-like, aromatic, or heterocyclic elements to test how rigidity and polarity affect degradation. Linker binding site selection and design can help identify attachment positions that preserve ligand recognition while enabling productive ternary-complex formation.
03Ternary Complex Cooperativity
Ternary-complex cooperativity describes how the binding of one protein partner affects recruitment of the other partner in the presence of a PROTAC. Positive cooperativity can improve degradation efficiency by stabilizing the target-PROTAC-E3 complex, while negative cooperativity may reduce productive complex formation. Because cooperativity depends on the combined surfaces of the target, ligase, and PROTAC, it cannot always be predicted from binary binding data.
Biophysical methods, structural modeling, and cellular degradation assays are often used together to understand ternary-complex behavior. BOC Sciences provides access to PROTAC ternary complex assay support for researchers who need to evaluate proximity-driven complex formation during degrader optimization.
04Managing the Hook Effect in PROTAC Studies
The hook effect is a concentration-dependent phenomenon in which excessive PROTAC levels can reduce ternary-complex formation by saturating the target protein and E3 ligase separately. Instead of forming a productive three-component complex, each protein may be occupied by separate PROTAC molecules, lowering degradation efficiency at high concentrations.
To identify the hook effect, researchers often examine broad concentration-response curves rather than relying on a single test concentration. A bell-shaped degradation profile may indicate that optimal ternary-complex formation occurs within a defined concentration window. Understanding this behavior helps teams select meaningful assay conditions and avoid misinterpreting a potent degrader as inactive due to inappropriate dosing ranges in research experiments.
PROTAC Products and Building Blocks
PROTAC research often requires access to multiple types of products, including finished degraders, E3 ligase ligand-linker conjugates, target protein ligands, functionalized linkers, and negative-control analogs. These materials help researchers build focused libraries, compare ligase recruitment strategies, validate degradation mechanisms, and accelerate the transition from concept to experimental testing.
PROTAC Degraders for Target Validation
Ready-to-use PROTAC degraders can support target validation, pathway analysis, and chemical knockdown studies when a relevant target and degrader pair is available. Researchers may use these compounds to compare degradation with inhibition, evaluate time-dependent target loss, and establish assay conditions before designing new analogs. BOC Sciences offers PROTAC products covering diverse target classes and degradation mechanisms for research use.
E3 ligase ligand-linker conjugates simplify PROTAC assembly by providing a prefunctionalized recruiter module ready for coupling with a target-binding ligand. These conjugates can reduce synthetic complexity in early library preparation and allow researchers to compare linkers while keeping the E3 ligand constant. BOC Sciences provides E3 ligase ligand-linker conjugate products for modular degrader construction.
Target protein ligands provide the recognition element that directs a PROTAC to the protein of interest. When selecting these ligands, researchers consider binding affinity, selectivity, chemical handles, and available structural information. In many projects, multiple target ligand analogs are evaluated to identify the most suitable exit vector and conjugation site for degrader design.
Functionalized PROTAC linkers enable systematic exploration of length, flexibility, polarity, and rigidity. A well-planned linker set can reveal whether degradation depends on a narrow geometric window or tolerates a broader range of molecular shapes. BOC Sciences provides linker products and linker design support for researchers building focused degrader libraries across target and ligase systems.
Advancing Your PROTAC Research?
BOC Sciences provides an extensive portfolio of in-stock PROTAC-related products to support targeted protein degradation studies. If the compound you need is not listed or you require a custom-designed molecule, our experienced team can deliver tailored synthesis solutions.
Request a Quote
PROTAC Assays and Degradation Evaluation
PROTAC evaluation usually requires a combined assay strategy because target binding, ternary-complex formation, ubiquitination, proteasome engagement, and selectivity may not correlate perfectly. A well-designed workflow helps researchers determine whether a degrader truly works through the ubiquitin-proteasome system and whether its activity is specific to the intended protein of interest.
In Vitro Biochemical Assessment
In vitro biochemical assays are used to confirm whether a PROTAC can engage both the protein of interest and the E3 ligase before entering cell-based evaluation.
- Ternary complex formation: Surface-based or homogeneous proximity assays can measure whether the PROTAC promotes POI-E3 association, with KD, cooperativity α, and complex stability as key readouts.
- In vitro ubiquitination: Reconstituted E1, E2, E3, ubiquitin, PROTAC, and target protein systems can be used to detect polyubiquitination signals, including high-molecular-weight target bands.
- Cell lysate degradation validation: Target-expressing cell lysates can be used with supplemented ubiquitination components to evaluate whether target protein reduction can be observed in a controlled biochemical environment.
Cell-Based Degradation and Activity Evaluation
Cell-based assays are central to PROTAC evaluation because they directly measure target protein loss in a relevant cellular setting.
- Western blot: Commonly used to quantify target protein reduction after compound treatment, with internal-control normalization supporting DC50 and Dmax analysis.
- High-content imaging: Useful when spatial information is important, such as monitoring nuclear proteins, membrane proteins, fluorescence intensity changes, or target localization shifts.
- Fluorescence or luminescence reporter systems: Reporter-based workflows can support higher-throughput degradation screening by linking target abundance to measurable optical signals.
- Live-cell kinetic monitoring: Real-time reporter systems can help track degradation kinetics continuously and support analysis of target reduction over time.
Mechanism Validation for UPS Dependence
Mechanism validation helps confirm that observed target reduction is mediated by the ubiquitin-proteasome system rather than an indirect pathway effect.
- Ubiquitination detection: The appearance of target protein ubiquitination signals in the presence of a PROTAC supports direct ubiquitin modification.
- Proteasome dependence: Loss or reduction of degradation under proteasome-pathway control conditions supports proteasome-dependent target processing.
- E3 ligase dependence: E3-related competition, inhibition, or genetic knockout studies can help confirm whether degradation requires the selected E3 ligase.
- Competitive ligand experiments: Excess E3 ligand can be used to test whether PROTAC activity depends on specific E3 recruitment.
Selectivity and Off-Target Degradation Profiling
Selectivity assessment determines whether a PROTAC mainly reduces the intended target or also affects additional proteins.
- Proteomics-based profiling: TMT or label-free quantification workflows can compare global protein abundance between control and PROTAC-treated samples to identify broad degradation patterns.
- Target-family comparison: Related proteins or paralogs can be monitored to understand whether degradation is selective within a protein family.
- Targeted Western blot confirmation: Potential off-target proteins identified through proteomics, target similarity, or E3 ligase biology can be verified with specific antibodies.
- Follow-up prioritization: Selectivity data can guide analog optimization, E3 ligase strategy adjustment, and further mechanism-focused studies.
Applications of PROTAC Technology
PROTAC technology is broadly used in chemical biology and drug discovery research to study protein function, validate targets, compare degradation with inhibition, and develop induced-proximity platforms. Because PROTACs act by reducing target protein abundance, they can reveal biology that is not accessible through activity blockade alone. This makes them useful tools for studying proteins with scaffolding roles, multi-domain functions, or complex pathway interactions.
Protein Function Validation
PROTACs can help researchers validate whether a protein is functionally important in a pathway by reducing its abundance and monitoring downstream effects. This approach may provide more complete insight than inhibition when a protein has non-catalytic roles or interacts with multiple partners. Degradation-based validation is commonly used in early discovery studies, pathway mapping, and target biology exploration.
Chemical Knockdown Research
Chemical knockdown using PROTACs enables rapid and reversible reduction of target protein levels in research systems. Compared with genetic knockdown methods, a degrader can provide temporal control and dose-dependent modulation. This allows researchers to examine acute target depletion, recovery after compound removal, and differences between partial and deep protein reduction.
Targeted Protein Degradation Platform Development
PROTACs are central to targeted protein degradation platform development. They provide a modular framework for testing target binders, E3 ligase recruiters, linkers, assay formats, and SAR strategies. BOC Sciences supports PROTAC degradation technology development for teams building or expanding degrader discovery workflows.
Discovery Programs for Difficult-to-Modulate Proteins
Many proteins are difficult to modulate with conventional occupancy-driven compounds because they lack a suitable active site or because their key biological roles are not limited to catalytic activity. PROTACs can provide an alternative research strategy when target removal is more informative than inhibition. Programs involving transcriptional regulators, epigenetic readers, kinases, scaffold proteins, and signaling adaptors often benefit from degradation-focused thinking.
PROTACs vs. Molecular Glues
PROTACs and molecular glues are two major induced-proximity strategies in targeted protein degradation research. Both approaches use cellular degradation machinery to reduce the abundance of selected proteins, but they differ substantially in molecular architecture, discovery logic, target recruitment, and optimization strategy. Understanding these differences helps research teams choose a suitable modality for degrader design, compound screening, mechanism studies, and product selection.
| Comparison Dimension | PROTACs | Molecular Glues |
| Structural Features | Bifunctional architecture containing a target protein ligand, a linker, and an E3 ligase ligand. | Monovalent small-molecule structure without a defined bifunctional linker. |
| Mechanism of Action | Physically brings the target protein and E3 ligase into proximity through a linker-mediated ternary complex. | Induces or stabilizes a protein-protein interaction between an E3 ligase and a recruited substrate protein. |
| Molecular Size | Usually larger and more complex because two ligands and a linker are combined in one molecule. | Usually more compact, which can simplify certain small-molecule design and handling considerations. |
| Design Strategy | Often supports rational and modular design by separately optimizing the target ligand, E3 ligand, and linker. | Often discovered through screening, phenotypic observation, or interface-focused optimization; rational design can be more challenging. |
| Target Scope | Typically requires a known ligand or binder for the protein of interest. | May enable exploration of proteins that lack a well-characterized small-molecule ligand. |
| Complex Formation | Ternary-complex geometry is strongly influenced by linker length, rigidity, polarity, and exit-vector selection. | Recruitment depends on protein surface complementarity and compound-induced interface remodeling. |
| Optimization Focus | Ligand pairing, linker chemistry, ternary-complex stability, degradation readouts, and structure-activity relationships. | Scaffold selection, neosubstrate recruitment, interface stabilization, selectivity mapping, and degradation profile interpretation. |
| Research Fit | Useful when target binders are available and modular degrader construction is feasible. | Useful when compact scaffolds or induced interface formation are central to the research hypothesis. |
Advantages and Limitations of PROTACs
PROTACs are highly valued in targeted protein degradation research because their modular architecture allows researchers to adjust each design element independently.
- Modular design logic: The target protein ligand, E3 ligase ligand, and linker can be modified separately to build focused analog series.
- Event-driven degradation concept: PROTACs are designed to reduce target protein abundance rather than only blocking a binding site.
- Lower occupancy dependence: Productive degradation may occur even when continuous target-site occupancy is not required in the same way as conventional inhibitors.
- Adjustable degradation profile: Linker length, linker composition, and E3 ligase selection can be tuned to explore different degradation outcomes.
However, PROTACs also present practical design challenges. Their larger molecular size, structural flexibility, synthesis complexity, and linker-dependent behavior can complicate compound optimization. In some systems, high compound concentrations may favor separate binary binding events rather than productive ternary-complex formation, leading to hook-related effects that require careful experimental interpretation.
Advantages and Limitations of Molecular Glues
Molecular glues offer a complementary approach because they are typically compact small molecules that induce proximity through interface remodeling rather than bifunctional bridging.
- Compact molecular architecture: Molecular glues often avoid the size and flexibility burden associated with long bifunctional degrader structures.
- Potential access to difficult targets: They may recruit proteins that do not have a known ligandable pocket suitable for conventional degrader construction.
- No linker optimization requirement: Because molecular glues generally lack a long linker, researchers can focus on scaffold refinement and interface recognition.
- Useful induced-proximity model: They provide a valuable route for studying how small molecules reshape protein-protein interactions.
The main challenge is discovery and predictability. Molecular glue activity can depend on subtle interface effects that are difficult to design rationally. Target prediction, selectivity interpretation, and degradation-spectrum control may require integrated screening, proteomics, structural analysis, and follow-up validation.
Why Choose BOC Sciences PROTAC Products?
BOC Sciences supports PROTAC research with product categories, design support, synthesis capabilities, assay resources, and technical communication aligned with the needs of B2B drug discovery and life science research teams. The goal is to help customers move from degrader concept to compound selection, experimental evaluation, and iterative optimization with practical scientific support.
 Broad Selection of PROTAC Degraders and Building Blocks
Broad Selection of PROTAC Degraders and Building Blocks
BOC Sciences offers PROTAC degraders, E3 ligase ligands, target protein ligands, ligand-linker conjugates, and linkers that support different stages of degrader research. This product breadth allows researchers to assemble focused libraries, compare ligase systems, and evaluate degradation strategies without relying on a single molecular design.
 Support for PROTAC Design and Custom Synthesis
Support for PROTAC Design and Custom Synthesis
For projects requiring molecules beyond catalog products, BOC Sciences can support custom degrader design and synthesis. Research teams can discuss target ligands, E3 recruiter options, linker strategies, conjugation handles, and analog series planning with technical specialists familiar with targeted protein degradation workflows.
 Technical Guidance for Product Selection
Technical Guidance for Product Selection
PROTAC product selection requires more than matching a target name. Researchers often need to consider mechanism, ligase recruitment, linker architecture, assay context, and compatibility with planned readouts. BOC Sciences provides practical technical communication to help customers select products and services that align with their research objectives.
 Integrated Resources for TPD Research Workflows
Integrated Resources for TPD Research Workflows
BOC Sciences connects chemistry, biology, and analysis resources across the targeted protein degradation workflow. From target ligand design and E3 ligase recruitment to degradation assays and SAR interpretation, customers can access integrated support that helps streamline PROTAC research planning and execution.
Client Feedback on BOC Sciences PROTAC Products
Useful Product Range for Degrader Screening
“The available PROTAC products and building blocks helped our research team compare several linker and E3 recruitment options during early degrader screening. The product information made it easier to narrow down a practical starting set for our assays.”
— Senior Scientist, US-based biotechnology research group
Clear Communication During Product Selection
“We needed guidance on selecting PROTAC linkers and E3 ligand-linker conjugates for a new target class. BOC Sciences provided responsive technical communication and helped us align product options with our planned degrader design workflow.”
— Discovery Chemistry Lead, European research organization
Support for Custom Analog Planning
“Our project required a small analog series with different linker lengths and conjugation handles. The discussion with BOC Sciences helped us organize a focused design plan and prioritize structures that were meaningful for SAR analysis.”
— Project Manager, North American drug discovery company
Helpful Resources for TPD Workflows
“BOC Sciences offered a practical combination of PROTAC products, assay-related resources, and technical support. This was helpful for coordinating chemistry and biology decisions within our targeted protein degradation research program.”
— Principal Investigator, UK-based life science institute
Frequently Asked Questions (FAQ)

Still have questions?
Contact Us
How do PROTACs degrade target proteins?
PROTACs degrade target proteins by bringing the selected protein into proximity with an E3 ubiquitin ligase. This induced-proximity event can support ubiquitin transfer to the target protein, after which the proteasome recognizes and degrades the ubiquitinated substrate. The process depends on target binding, E3 ligase recruitment, ternary-complex geometry, linker design, and cellular assay context, so degradation performance must be confirmed with functional readouts rather than predicted only from binding affinity.
How should researchers choose PROTAC linkers?
Researchers should choose PROTAC linkers by considering linker length, flexibility, polarity, rigidity, attachment points, and compatibility with both ligands. The linker must preserve target and E3 ligase binding while positioning the two proteins in a geometry that supports productive ubiquitination. BOC Sciences can support linker selection through PROTAC linker products, linker design resources, and custom analog planning for research teams comparing multiple degrader architectures.
What E3 ligases are common in PROTAC design?
Common E3 ligases used in PROTAC design include cereblon, VHL, IAP family members, and MDM2-related systems. Each ligase system has different ligand chemistry, expression context, localization behavior, and ternary-complex requirements. Because degradation depends on the compatibility between the target protein and recruited ligase, researchers often compare several E3 recruitment strategies before selecting the most informative degrader series for further study.
What assays help evaluate PROTAC activity?
PROTAC activity is commonly evaluated through target degradation analysis, binding affinity measurement, ternary-complex assessment, ubiquitination studies, and proteasome-pathway controls. Concentration-response and time-course experiments are also useful for identifying degradation kinetics, selectivity trends, concentration windows, and possible hook-effect behavior. BOC Sciences provides assay-related support that can help researchers connect compound structure with target protein loss, pathway dependence, and structure-activity relationship interpretation.
Can BOC Sciences support custom PROTAC projects?
BOC Sciences can support custom PROTAC projects involving target ligand selection, E3 ligase ligand choice, linker design, degrader synthesis, analog series planning, and degradation-focused evaluation. This support is designed for research use and can help teams move from a target hypothesis to a practical set of compounds for experimental testing. Technical communication can also help align product selection with assay design and project objectives.
Discover More Research Products
Explore featured products that can expand your research options and accelerate your next discovery.
Expert Services to Move Your Project Forward
Access end-to-end service solutions that help bring efficiency, flexibility, and expertise to your research pipeline.
Insights and Resources
News
Technical Information

![(2S,4R)-N-((S)-2-(tert-Butyl)-17-((S)-4-(4-chlorophenyl)-2,3,9-trimethyl-6H-thieno[3,2-f][1,2,4]triazolo[4,3-a][1,4]diazepin-6-yl)-4,16-dioxo-6,9,12-trioxa-3,15-diazaheptadecan-1-oyl)-4-hydroxy-1-(4-(4-methylthiazol-5-yl)benzyl)pyrrolidine-2-carboxamide](https://resource.bocsci.com/structure/1797406-69-9.gif)



















 Fig.1 PROTAC-mediated targeted protein degradation workflow (BOC Sciences Original).
Fig.1 PROTAC-mediated targeted protein degradation workflow (BOC Sciences Original).